Outline the process of prokaryotic transcription and translation
The prokaryotes, which include bacteria and archaea, are mostly single-celled organisms that, by definition, lack membrane-bound nuclei and other organelles. A bacterial chromosome is a covalently closed circle that, unlike eukaryotic chromosomes, is not organized around histone proteins. The central region of the cell in which prokaryotic DNA resides is called the nucleoid. In addition, prokaryotes often have abundant plasmids, which are shorter circular DNA molecules that may only contain one or a few genes. Plasmids can be transferred independently of the bacterial chromosome during cell division and often carry traits such as antibiotic resistance. Because of these unique features, transcription and gene regulation is somewhat different between prokaryotic cells and eukaryotic ones.
Learning Objectives
- Understand the basic steps in the transcription of DNA into RNA in prokaryotic cells
- Understand the basics of prokaryotic translation and how it differs from eukaryotic translation
Prokaryotic Transcription
Initiation of Transcription in Prokaryotes
Prokaryotes do not have membrane-enclosed nuclei. Therefore, the processes of transcription, translation, and mRNA degradation can all occur simultaneously. The intracellular level of a bacterial protein can quickly be amplified by multiple transcription and translation events occurring concurrently on the same DNA template. Prokaryotic transcription often covers more than one gene and produces polycistronic mRNAs that specify more than one protein.
Our discussion here will exemplify transcription by describing this process in Escherichia coli, a well-studied bacterial species. Although some differences exist between transcription in E. coli and transcription in archaea, an understanding of E. coli transcription can be applied to virtually all bacterial species.
Prokaryotic RNA Polymerase
Prokaryotes use the same RNA polymerase to transcribe all of their genes. In E. coli, the polymerase is composed of five polypeptide subunits, two of which are identical. Four of these subunits, denoted α, α, β, and β′ comprise the polymerase core enzyme. These subunits assemble every time a gene is transcribed, and they disassemble once transcription is complete. Each subunit has a unique role; the two α-subunits are necessary to assemble the polymerase on the DNA; the β-subunit binds to the ribonucleoside triphosphate that will become part of the nascent “recently born” mRNA molecule; and the β′ binds the DNA template strand. The fifth subunit, σ, is involved only in transcription initiation. It confers transcriptional specificity such that the polymerase begins to synthesize mRNA from an appropriate initiation site. Without σ, the core enzyme would transcribe from random sites and would produce mRNA molecules that specified protein gibberish. The polymerase comprised of all five subunits is called the holoenzyme.
Prokaryotic Promoters

Figure 1. The σ subunit of prokaryotic RNA polymerase recognizes consensus sequences found in the promoter region upstream of the transcription start sight. The σ subunit dissociates from the polymerase after transcription has been initiated.
A promoter is a DNA sequence onto which the transcription machinery binds and initiates transcription. In most cases, promoters exist upstream of the genes they regulate. The specific sequence of a promoter is very important because it determines whether the corresponding gene is transcribed all the time, some of the time, or infrequently. Although promoters vary among prokaryotic genomes, a few elements are conserved. At the -10 and -35 regions upstream of the initiation site, there are two promoter consensus sequences, or regions that are similar across all promoters and across various bacterial species (Figure 1).
The -10 consensus sequence, called the -10 region, is TATAAT. The -35 sequence, TTGACA, is recognized and bound by σ. Once this interaction is made, the subunits of the core enzyme bind to the site. The A–T-rich -10 region facilitates unwinding of the DNA template, and several phosphodiester bonds are made. The transcription initiation phase ends with the production of abortive transcripts, which are polymers of approximately 10 nucleotides that are made and released.
Elongation and Termination in Prokaryotes
The transcription elongation phase begins with the release of the σ subunit from the polymerase. The dissociation of σ allows the core enzyme to proceed along the DNA template, synthesizing mRNA in the 5′ to 3′ direction at a rate of approximately 40 nucleotides per second. As elongation proceeds, the DNA is continuously unwound ahead of the core enzyme and rewound behind it (Figure 2). The base pairing between DNA and RNA is not stable enough to maintain the stability of the mRNA synthesis components. Instead, the RNA polymerase acts as a stable linker between the DNA template and the nascent RNA strands to ensure that elongation is not interrupted prematurely.

Figure 2. Click for a larger image. During elongation, the prokaryotic RNA polymerase tracks along the DNA template, synthesizes mRNA in the 5′ to 3′ direction, and unwinds and rewinds the DNA as it is read.
Prokaryotic Termination Signals
Once a gene is transcribed, the prokaryotic polymerase needs to be instructed to dissociate from the DNA template and liberate the newly made mRNA. Depending on the gene being transcribed, there are two kinds of termination signals. One is protein-based and the other is RNA-based. Rho-dependent termination is controlled by the rho protein, which tracks along behind the polymerase on the growing mRNA chain. Near the end of the gene, the polymerase encounters a run of G nucleotides on the DNA template and it stalls. As a result, the rho protein collides with the polymerase. The interaction with rho releases the mRNA from the transcription bubble.
Rho-independent termination is controlled by specific sequences in the DNA template strand. As the polymerase nears the end of the gene being transcribed, it encounters a region rich in C–G nucleotides. The mRNA folds back on itself, and the complementary C–G nucleotides bind together. The result is a stable hairpin that causes the polymerase to stall as soon as it begins to transcribe a region rich in A–T nucleotides. The complementary U–A region of the mRNA transcript forms only a weak interaction with the template DNA. This, coupled with the stalled polymerase, induces enough instability for the core enzyme to break away and liberate the new mRNA transcript.
Upon termination, the process of transcription is complete. By the time termination occurs, the prokaryotic transcript would already have been used to begin synthesis of numerous copies of the encoded protein because these processes can occur concurrently. The unification of transcription, translation, and even mRNA degradation is possible because all of these processes occur in the same 5′ to 3′ direction, and because there is no membranous compartmentalization in the prokaryotic cell (Figure 3). In contrast, the presence of a nucleus in eukaryotic cells precludes simultaneous transcription and translation.

Figure 3. Multiple polymerases can transcribe a single bacterial gene while numerous ribosomes concurrently translate the mRNA transcripts into polypeptides. In this way, a specific protein can rapidly reach a high concentration in the bacterial cell.
Visit this BioStudio animation to see the process of prokaryotic transcription.
Practice Questions
Which subunit of the E. coli polymerase confers specificity to transcription?
- α
- β
- β′
- σ
The -10 and -35 regions of prokaryotic promoters are called consensus sequences because ________.
- they are identical in all bacterial species
- they are similar in all bacterial species
- they exist in all organisms
- they have the same function in all organisms
Prokaryotic Translation
Translation is similar in prokaryotes and eukaryotes. Here we will explore how translation occurs in E. coli, a representative prokaryote, and specify any differences between bacterial and eukaryotic translation.
Initiation
The initiation of protein synthesis begins with the formation of an initiation complex. In E. coli, this complex involves the small 30S ribosome, the mRNA template, three initiation factors that help the ribosome assemble correctly, guanosine triphosphate (GTP) that acts as an energy source, and a special initiator tRNA carrying N-formyl-methionine (fMet-tRNAfMet) (Figure 4). The initiator tRNA interacts with the start codon AUG of the mRNA and carries a formylated methionine (fMet). Because of its involvement in initiation, fMet is inserted at the beginning (N terminus) of every polypeptide chain synthesized by E. coli. In E. coli mRNA, a leader sequence upstream of the first AUG codon, called the Shine-Dalgarno sequence (also known as the ribosomal binding site AGGAGG), interacts through complementary base pairing with the rRNA molecules that compose the ribosome. This interaction anchors the 30S ribosomal subunit at the correct location on the mRNA template. At this point, the 50S ribosomal subunit then binds to the initiation complex, forming an intact ribosome.
In eukaryotes, initiation complex formation is similar, with the following differences:
- The initiator tRNA is a different specialized tRNA carrying methionine, called Met-tRNAi
- Instead of binding to the mRNA at the Shine-Dalgarno sequence, the eukaryotic initiation complex recognizes the 5′ cap of the eukaryotic mRNA, then tracks along the mRNA in the 5′ to 3′ direction until the AUG start codon is recognized. At this point, the 60S subunit binds to the complex of Met-tRNAi, mRNA, and the 40S subunit.

Figure 4. Translation in bacteria begins with the formation of the initiation complex, which includes the small ribosomal subunit, the mRNA, the initiator tRNA carrying N-formyl-methionine, and initiation factors. Then the 50S subunit binds, forming an intact ribosome.
Elongation
In prokaryotes and eukaryotes, the basics of elongation of translation are the same. In E. coli, the binding of the 50S ribosomal subunit to produce the intact ribosome forms three functionally important ribosomal sites: The A (aminoacyl) site binds incoming charged aminoacyl tRNAs. The P (peptidyl) site binds charged tRNAs carrying amino acids that have formed peptide bonds with the growing polypeptide chain but have not yet dissociated from their corresponding tRNA. The E (exit) site releases dissociated tRNAs so that they can be recharged with free amino acids. There is one notable exception to this assembly line of tRNAs: During initiation complex formation, bacterial fMet−tRNAfMet or eukaryotic Met-tRNAi enters the P site directly without first entering the A site, providing a free A site ready to accept the tRNA corresponding to the first codon after the AUG.
Elongation proceeds with single-codon movements of the ribosome each called a translocation event. During each translocation event, the charged tRNAs enter at the A site, then shift to the P site, and then finally to the E site for removal. Ribosomal movements, or steps, are induced by conformational changes that advance the ribosome by three bases in the 3′ direction. Peptide bonds form between the amino group of the amino acid attached to the A-site tRNA and the carboxyl group of the amino acid attached to the P-site tRNA. The formation of each peptide bond is catalyzed by peptidyl transferase, an RNA-based ribozyme that is integrated into the 50S ribosomal subunit. The amino acid bound to the P-site tRNA is also linked to the growing polypeptide chain. As the ribosome steps across the mRNA, the former P-site tRNA enters the E site, detaches from the amino acid, and is expelled. Several of the steps during elongation, including binding of a charged aminoacyl tRNA to the A site and translocation, require energy derived from GTP hydrolysis, which is catalyzed by specific elongation factors. Amazingly, the E. coli translation apparatus takes only 0.05 seconds to add each amino acid, meaning that a 200 amino-acid protein can be translated in just 10 seconds.
Termination
The termination of translation occurs when a nonsense codon (UAA, UAG, or UGA) is encountered for which there is no complementary tRNA. On aligning with the A site, these nonsense codons are recognized by release factors in prokaryotes and eukaryotes that result in the P-site amino acid detaching from its tRNA, releasing the newly made polypeptide. The small and large ribosomal subunits dissociate from the mRNA and from each other; they are recruited almost immediately into another translation init iation complex.
In summary, there are several key features that distinguish prokaryotic gene expression from that seen in eukaryotes. These are illustrated in Figure 5 and listed in Table 1.
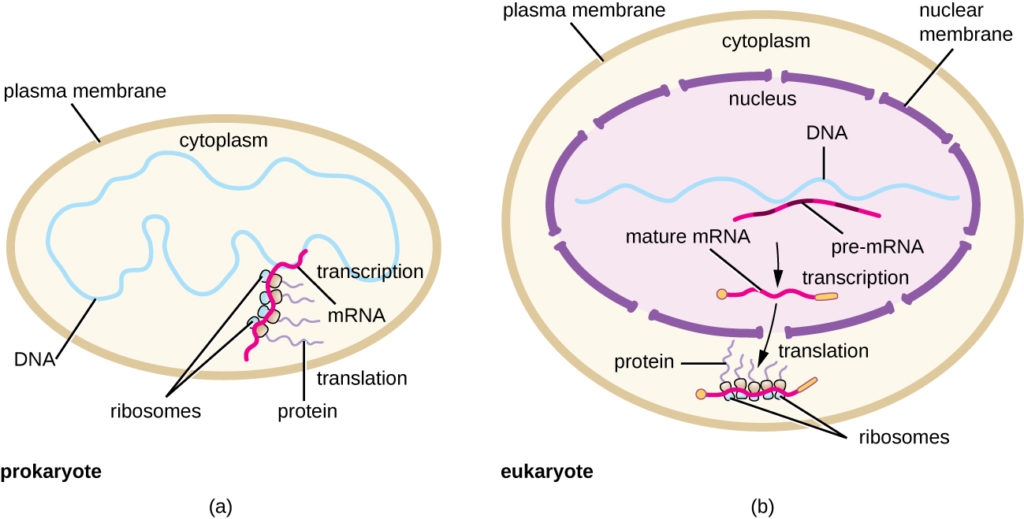
Figure 5. (a) In prokaryotes, the processes of transcription and translation occur simultaneously in the cytoplasm, allowing for a rapid cellular response to an environmental cue. (b) In eukaryotes, transcription is localized to the nucleus and translation is localized to the cytoplasm, separating these processes and necessitating RNA processing for stability.
| Table 1. Comparison of Translation in Bacteria Versus Eukaryotes | ||
|---|---|---|
| Property | Bacteria | Eukaryotes |
| Ribosomes | 70S
|
80S
|
| Amino acid carried by initiator tRNA | fMet | Met |
| Shine-Dalgarno sequence in mRNA | Present | Absent |
| Simultaneous transcription and translation | Yes | No |
Check Your Understanding
Answer the question(s) below to see how well you understand the topics covered in the previous section. This short quiz does not count toward your grade in the class, and you can retake it an unlimited number of times.
Use this quiz to check your understanding and decide whether to (1) study the previous section further or (2) move on to the next section.
Candela Citations
- Introduction to Prokaryotic Transcription and Translation. Authored by: Shelli Carter and Lumen Learning. Provided by: Lumen Learning. License: CC BY: Attribution
- Biology. Provided by: OpenStax CNX. Located at: http://cnx.org/contents/185cbf87-c72e-48f5-b51e-f14f21b5eabd@10.8. License: CC BY: Attribution. License Terms: Download for free at http://cnx.org/contents/185cbf87-c72e-48f5-b51e-f14f21b5eabd@10.8