Learning Outcomes
- Understand the basics of prokaryotic translation and how it differs from eukaryotic translation
As with mRNA synthesis, protein synthesis can be divided into three phases: initiation, elongation, and termination. The process of translation is similar in prokaryotes and eukaryotes. Here we’ll explore how translation occurs in E. coli, a representative prokaryote, and specify any differences between prokaryotic and eukaryotic translation.
Initiation
Protein synthesis begins with the formation of an initiation complex. In E. coli, this complex involves the small 30S ribosome, the mRNA template, three initiation factors (IFs; IF-1, IF-2, and IF-3), and a special initiator tRNA, called tRNAMetf.
In E. coli mRNA, a sequence upstream of the first AUG codon, called the Shine-Dalgarno sequence (AGGAGG), interacts with the rRNA molecules that compose the ribosome. This interaction anchors the 30S ribosomal subunit at the correct location on the mRNA template. Guanosine triphosphate (GTP), which is a purine nucleotide triphosphate, acts as an energy source during translation—both at the start of elongation and during the ribosome’s translocation. Binding of the mRNA to the 30S ribosome also requires IF-III.
The initiator tRNA then interacts with the start codon AUG (or rarely, GUG). This tRNA carries the amino acid methionine, which is formylated after its attachment to the tRNA. The formylation creates a “faux” peptide bond between the formyl carboxyl group and the amino group of the methionine. Binding of the fMet-tRNAMetf is mediated by the initiation factor IF-2. The fMet begins every polypeptide chain synthesized by E. coli, but it is usually removed after translation is complete. When an in-frame AUG is encountered during translation elongation, a non-formylated methionine is inserted by a regular Met-tRNAMet. After the formation of the initiation complex, the 30S ribosomal subunit is joined by the 50S subunit to form the translation complex. In eukaryotes, a similar initiation complex forms, comprising mRNA, the 40S small ribosomal subunit, eukaryotic IFs, and nucleoside triphosphates (GTP and ATP). The methionine on the charged initiator tRNA, called Met-tRNAi, is not formylated. However, Met-tRNAi is distinct from other Met-tRNAs in that it can bind IFs.
Instead of depositing at the Shine-Dalgarno sequence, the eukaryotic initiation complex recognizes the 7-methylguanosine cap at the 5′ end of the mRNA. A cap-binding protein (CBP) and several other IFs assist the movement of the ribosome to the 5′ cap. Once at the cap, the initiation complex tracks along the mRNA in the 5′ to 3′ direction, searching for the AUG start codon. Many eukaryotic mRNAs are translated from the first AUG, but this is not always the case. According to Kozak’s rules, the nucleotides around the AUG indicate whether it is the correct start codon. Kozak’s rules state that the following consensus sequence must appear around the AUG of vertebrate genes: 5′-gccRccAUGG-3′. The R (for purine) indicates a site that can be either A or G, but cannot be C or U. Essentially, the closer the sequence is to this consensus, the higher the efficiency of translation.
Once the appropriate AUG is identified, the other proteins and CBP dissociate, and the 60S subunit binds to the complex of Met-tRNAi, mRNA, and the 40S subunit. This step completes the initiation of translation in eukaryotes.
In eukaryotes, initiation complex formation is similar, with the following differences:
- The initiator tRNA is a different specialized tRNA carrying methionine, called Met-tRNAi
- Instead of binding to the mRNA at the Shine-Dalgarno sequence, the eukaryotic initiation complex recognizes the 5′ cap of the eukaryotic mRNA, then tracks along the mRNA in the 5′ to 3′ direction until the AUG start codon is recognized. At this point, the 60S subunit binds to the complex of Met-tRNAi, mRNA, and the 40S subunit.

Figure 1. Translation in bacteria begins with the formation of the initiation complex, which includes the small ribosomal subunit, the mRNA, the initiator tRNA carrying N-formyl-methionine, and initiation factors. Then the 50S subunit binds, forming an intact ribosome.
Elongation
Figure 2. Translation begins when an initiator tRNA anticodon recognizes a start codon on mRNA bound to a small ribosomal subunit. The large ribosomal subunit joins the small subunit, and a second tRNA is recruited. As the mRNA moves relative to the ribosome, successive tRNAs move through the ribosome and the polypeptide chain is formed. Entry of a release factor into the A site terminates translation and the components dissociate.
In prokaryotes and eukaryotes, the basics of elongation are the same, so we will review elongation from the perspective of E. coli. When the translation complex is formed, the tRNA binding region of the ribosome consists of three compartments. The A (aminoacyl) site binds incoming charged aminoacyl tRNAs. The P (peptidyl) site binds charged tRNAs carrying amino acids that have formed peptide bonds with the growing polypeptide chain but have not yet dissociated from their corresponding tRNA. The E (exit) site releases dissociated tRNAs so that they can be recharged with free amino acids. The initiating methionyl-tRNA, however, occupies the P site at the beginning of the elongation phase of translation in both prokaryotes and eukaryotes.
During translation elongation, the mRNA template provides tRNA binding specificity. As the ribosome moves along the mRNA, each mRNA codon comes into register, and specific binding with the corresponding charged tRNA anticodon is ensured. If mRNA were not present in the elongation complex, the ribosome would bind tRNAs nonspecifically and randomly.
Elongation proceeds with charged tRNAs sequentially entering and leaving the ribosome as each new amino acid is added to the polypeptide chain. Movement of a tRNA from A to P to E site is induced by conformational changes that advance the ribosome by three bases in the 3′ direction. The energy for each step along the ribosome is donated by elongation factors that hydrolyze GTP. GTP energy is required both for the binding of a new aminoacyl-tRNA to the A site and for its translocation to the P site after formation of the peptide bond. Peptide bonds form between the amino group of the amino acid attached to the A-site tRNA and the carboxyl group of the amino acid attached to the P-site tRNA. The formation of each peptide bond is catalyzed by peptidyl transferase, an RNA-based enzyme that is integrated into the 50S ribosomal subunit. The energy for each peptide bond formation is derived from the high-energy bond linking each amino acid to its tRNA. After peptide bond formation, the A-site tRNA that now holds the growing peptide chain moves to the P site, and the P-site tRNA that is now empty moves to the E site and is expelled from the ribosome (Figure 2). Amazingly, the E. coli translation apparatus takes only 0.05 seconds to add each amino acid, meaning that a 200-amino-acid protein can be translated in just 10 seconds.
Termination
Termination of translation occurs when a nonsense codon (UAA, UAG, or UGA) is encountered. Upon aligning with the A site, these nonsense codons are recognized by protein release factors that resemble tRNAs. The releasing factors in both prokaryotes and eukaryotes instruct peptidyl transferase to add a water molecule to the carboxyl end of the P-site amino acid. This reaction forces the P-site amino acid to detach from its tRNA, and the newly made protein is released. The small and large ribosomal subunits dissociate from the mRNA and from each other; they are recruited almost immediately into another translation initiation complex. After many ribosomes have completed translation, the mRNA is degraded so the nucleotides can be reused in another transcription reaction.
In summary, there are several key features that distinguish prokaryotic gene expression from that seen in eukaryotes. These are illustrated in Figure 3 and listed in Table 1.
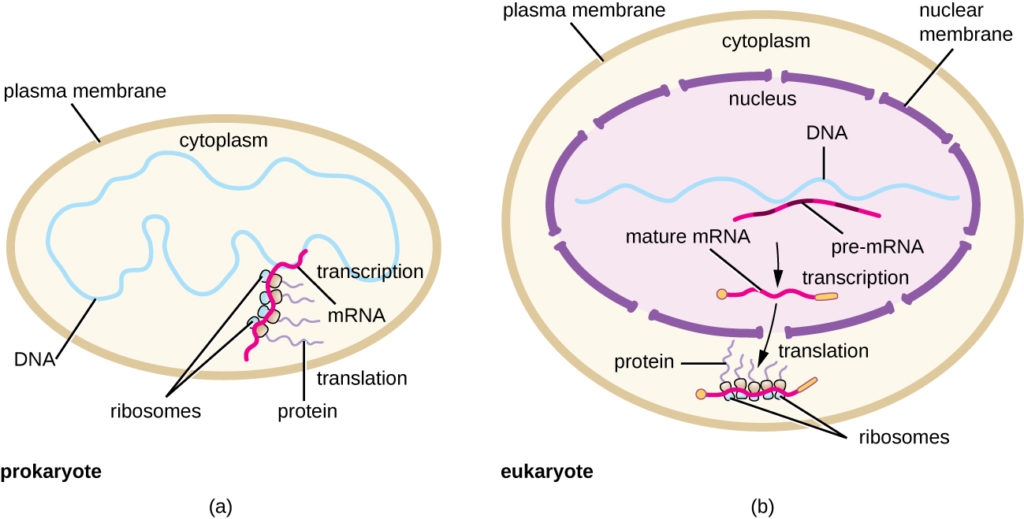
Figure 3. (a) In prokaryotes, the processes of transcription and translation occur simultaneously in the cytoplasm, allowing for a rapid cellular response to an environmental cue. (b) In eukaryotes, transcription is localized to the nucleus and translation is localized to the cytoplasm, separating these processes and necessitating RNA processing for stability.
| Table 1. Comparison of Translation in Bacteria Versus Eukaryotes | ||
|---|---|---|
| Property | Bacteria | Eukaryotes |
| Ribosomes | 70S
|
80S
|
| Amino acid carried by initiator tRNA | fMet | Met |
| Shine-Dalgarno sequence in mRNA | Present | Absent |
| Simultaneous transcription and translation | Yes | No |
Try It
Candela Citations
- Biology 2e. Provided by: OpenStax. Located at: http://cnx.org/contents/185cbf87-c72e-48f5-b51e-f14f21b5eabd@10.8. License: CC BY: Attribution. License Terms: Access for free at https://openstax.org/books/biology-2e/pages/1-introduction
