Learning Outcomes
- Explain the process of epigenetic regulation
The human genome encodes over 20,000 genes; each of the 23 pairs of human chromosomes encodes thousands of genes. The DNA in the nucleus is precisely wound, folded, and compacted into chromosomes so that it will fit into the nucleus. It is also organized so that specific segments can be accessed as needed by a specific cell type.
The first level of organization, or packing, is the winding of DNA strands around histone proteins. Histones package and order DNA into structural units called nucleosome complexes, which can control the access of proteins to the DNA regions (Figure 1a). Under the electron microscope, this winding of DNA around histone proteins to form nucleosomes looks like small beads on a string (Figure 1b). These beads (histone proteins) can move along the string (DNA) and change the structure of the molecule.
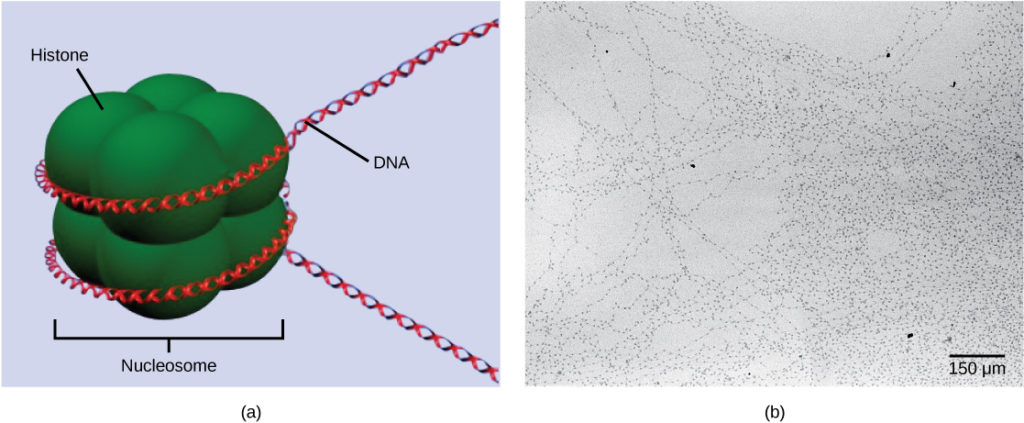
Figure 1. DNA is folded around histone proteins to create (a) nucleosome complexes. These nucleosomes control the access of proteins to the underlying DNA. When viewed through an electron microscope (b), the nucleosomes look like beads on a string. (credit “micrograph”: modification of work by Chris Woodcock)
If DNA encoding a specific gene is to be transcribed into RNA, the nucleosomes surrounding that region of DNA can slide down the DNA to open that specific chromosomal region and allow for the transcriptional machinery (RNA polymerase) to initiate transcription (Figure 2). Nucleosomes can move to open the chromosome structure to expose a segment of DNA, but do so in a very controlled manner.
Practice Question
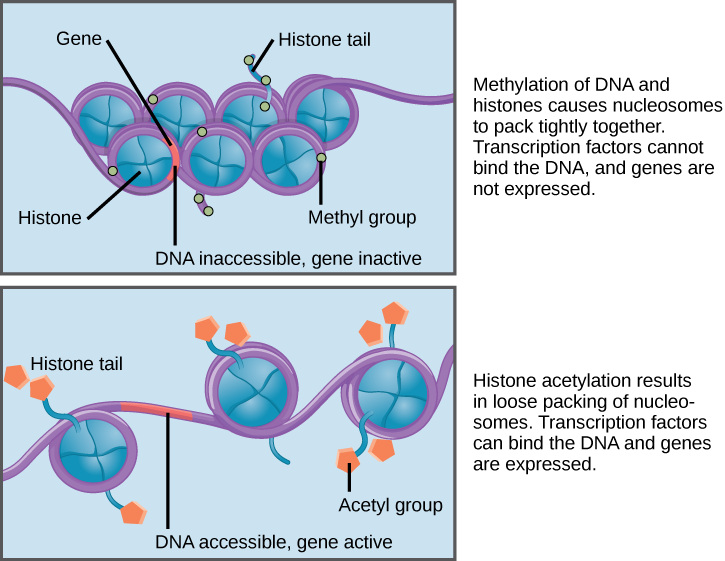
Figure 2. Nucleosomes can slide along DNA. When nucleosomes are spaced closely together (top), transcription factors cannot bind and gene expression is turned off. When the nucleosomes are spaced far apart (bottom), the DNA is exposed. Transcription factors can bind, allowing gene expression to occur. Modifications to the histones and DNA affect nucleosome spacing.
In people who have two X chromosomes, one of the two X chromosomes is inactivated during embryonic development because of epigenetic changes to the chromatin. What impact do you think these changes would have on nucleosome packing?
How closely the histone proteins associate with the DNA is regulated by signals found on both the histone proteins and on the DNA. These signals are functional groups added to histone proteins or to DNA and determine whether a chromosomal region should be open or closed (Figure 3 depicts modifications to histone proteins and DNA). These tags are not permanent, but may be added or removed as needed. Some chemical groups (phosphate, methyl, or acetyl groups) are attached to specific amino acids in histone “tails” at the N-terminus of the protein. These groups do not alter the DNA base sequence, but they do alter how tightly wound the DNA is around the histone proteins. DNA is a negatively charged molecule and unmodified histones are positively charged; therefore, changes in the charge of the histone will change how tightly wound the DNA molecule will be. By adding chemical modifications like acetyl groups, the charge becomes less positive, and the binding of DNA to the histones is relaxed. Altering the location of nucleosomes and the tightness of histone binding opens some regions of chromatin to transcription and closes others.
The DNA molecule itself can also be modified by methylation. DNA methylation occurs within very specific regions called CpG islands. These are stretches with a high frequency of cytosine and guanine dinucleotide DNA pairs (CG) found in the promoter regions of genes. The cytosine member of the CG pair can be methylated (a methyl group is added). Methylated genes are usually silenced, although methylation may have other regulatory effects. In some cases, genes that are silenced during the development of the gametes of one parent are transmitted in their silenced condition to the offspring. Such genes are said to be imprinted. Parental diet or other environmental conditions may also affect the methylation patterns of genes, which in turn modifies gene expression. Changes in chromatin organization interact with DNA methylation. DNA methyltransferases appear to be attracted to chromatin regions with specific histone modifications. Highly methylated (hypermethylated) DNA regions with deacetylated histones are tightly coiled and transcriptionally inactive.

Figure 3. Histone proteins and DNA nucleotides can be modified chemically. Modifications affect nucleosome spacing and gene expression. (credit: modification of work by NIH)
Epigenetic changes are not permanent, although they often persist through multiple rounds of cell division and may even cross generational lines. Chromatin remodeling alters the chromosomal structure (open or closed) as needed. If a gene is to be transcribed, the histone proteins and DNA in the chromosomal region encoding that gene are modified in a way that opens the promoter region to allow RNA polymerase and other proteins, called transcription factors, to bind and initiate transcription. If a gene is to remain turned off, or silenced, the histone proteins and DNA have different modifications that signal a closed chromosomal configuration. In this closed configuration, the RNA polymerase and transcription factors do not have access to the DNA and transcription cannot occur (Figure 3).
View this video that describes how epigenetic regulation controls gene expression.
In Summary: Eukaryotic Epigenetic Gene Regulation
In eukaryotic cells, the first stage of gene expression control occurs at the epigenetic level. Epigenetic mechanisms control access to the chromosomal region to allow genes to be turned on or off. These mechanisms control how DNA is packed into the nucleus by regulating how tightly the DNA is wound around histone proteins. The addition or removal of chemical modifications (or flags) to histone proteins or DNA signals to the cell to open or close a chromosomal region. Therefore, eukaryotic cells can control whether a gene is expressed by controlling accessibility to transcription factors and the binding of RNA polymerase to initiate transcription.
Try It
Candela Citations
- Biology 2e. Provided by: OpenStax. Located at: http://cnx.org/contents/185cbf87-c72e-48f5-b51e-f14f21b5eabd@10.8. License: CC BY: Attribution. License Terms: Access for free at https://openstax.org/books/biology-2e/pages/1-introduction
